Indication Expansion
Indication expansion, finding new indications for an existing therapeutic target, is a common approach in drug development. Although seen as a risk mitigating strategy, selecting the most appropriate indication to pursue raises many difficult challenges including:
Euretos addresses these challenges through a unique combination of novel predictions from computational disease models coupled with the ability for domain experts to assess these independently in the Euretos AI platform.
Euretos approach to indication selection
Based on its many collaborations with leading pharma and biotech companies on indication selection, Euretos has developed unique, AI-powered computational disease models that predict truly novel indication candidates for a target by:
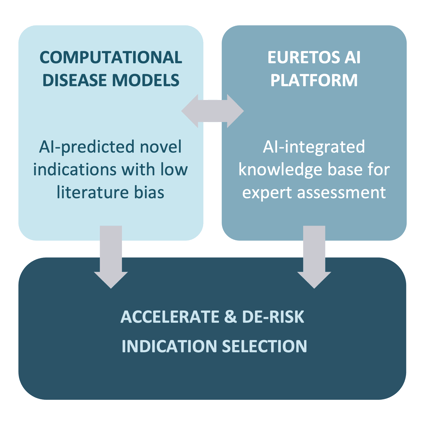
We know from many previous projects that the ability to assess predicted indications by target or disease experts is crucial to move a candidate forward in the selection process. We therefore empower researchers to independently evaluate predicted indications through:
Having proven computational disease models and the target assessment environment already in place, we can significantly accelerate and de-risk the process of indication discovery and assessment.
Predicting novel indications through computational disease models
Euretos has created computational disease models that have been tried and tested in many collaborations with leading pharma and biotech companies.
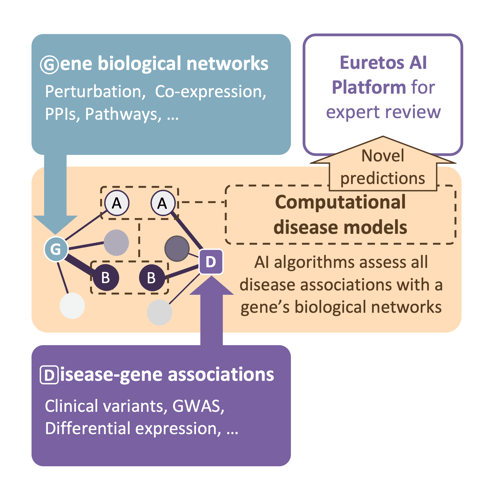
We predict novel indications by assessing whether a target’s biological interactors are linked to known disease associations. In particular we assess:
For each target, we use machine learning to evaluate for thousands of indications the potential associations the gene’s multiple biological networks may have with each disease. We combine the weighted predictions from these multiple algorithms into a single gene-disease score.
The end result of this is a ranked list of indications purely based on biological data and known disease indications, minimising potential literature bias.
Empower researchers to independently evaluate predicted indications
The ability to assess the AI-predicted indications by target or disease experts is crucial to move a candidate forward in the selection process. For over 10 years, Euretos has been developing the Euretos AI Platform which offers a one-of-a-kind biomolecular context for target exploration & assessment.
The Euretos knowledge base uses AI technologies, such as machine reading, to harmonise knowledge from biomolecular databases, publications and patents. This includes biological interaction networks, gene expression in tissues, cell types, subcellular location and tumors, pathway interactions, regulatory and signaling responses, clinical variant associations to disease and phenotypes, clinical trial outcomes, target tractability and patents.
The Euretos AI Platform consists of multiple applications that empower researchers to draw their own, evidence-based conclusions providing them with:
Results
The unique combination of novel predictions from the Euretos computational disease models with the ability for domain experts to assess these independently in the Euretos AI platform is very effective. Pharma and biotech companies are able to build within a very short time frame the confidence necessary to decide which indications to move forward for further development.







Keep up with our latest news and events. Sign up for our newsletter.